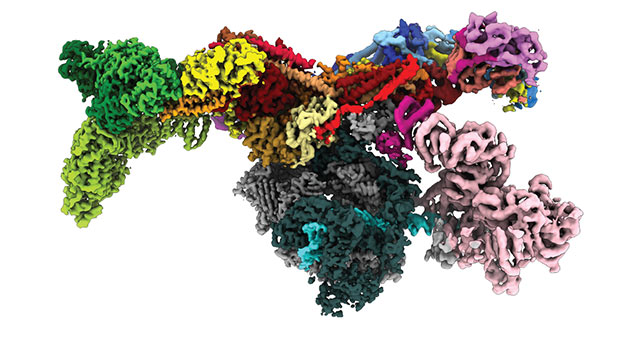
For the first time ever, a Northwestern University-led research team has peered inside a human cell to view a multi-subunit machine responsible for regulating gene expression.
Called the Mediator-bound pre-initiation complex (Med-PIC), the structure is a key player in determining which genes are activated and which are suppressed. Mediator helps position the rest of the complex — RNA polymerase II and the general transcription factors — at the beginning of genes that the cell wants to transcribe.
The researchers visualized the complex in high resolution using cryogenic electron microscopy (cryo-EM), enabling them to better understand how it works. Because this complex plays a role in many diseases, including cancer, neurodegenerative diseases, HIV and metabolic disorders, researchers’ new understanding of its structure could potentially be leveraged to treat disease.

“This machine is so basic to every branch of modern molecular biology in the context of gene expression,” said Northwestern’s Yuan He, senior author of the study. “Visualizing the structure in 3D will help us answer basic biological questions, such as how DNA is copied to RNA.”
“Seeing this structure allows us to understand how it works,” added Ryan Abdella, the paper’s co-first author. “It’s like taking apart a common household appliance to see how everything fits together. Now we can understand how the proteins in the complex come together to perform their function.”
The study will be published March 11 in the journal Science. This marks the first time the human Mediator complex has been visualized in 3D in the human cell.
He is an assistant professor of molecular biosciences in Northwestern’s Weinberg College of Arts and Sciences and member of the Chemistry of Life Processes Institute. Abdella and Anna Talyzina, both graduate students in the He lab, are co-first authors of the paper.
Famed biochemist Roger Kornberg discovered the Mediator complex in yeast in 1990, a project for which he won the 2006 Nobel Prize in Chemistry. But Mediator comprises a daunting 26 subunits — 56 total when combined with the pre-initiation complex — it’s taken researchers until now to obtain high-resolution images of the human version.
“It’s a technically quite challenging project,” He said. “These complexes are scarce. It takes hundreds of liters of human cells, which are very hard to grow, to obtain small amounts of the protein complexes.”
A breakthrough came when He’s team put the sample on a single layer of graphene oxide. By providing this support, the graphene sheet minimized the amount of sample needed for imaging. And compared to the typical support used — amorphous carbon — graphene improved the signal-to-noise ratio for higher-resolution imaging.
After preparing the sample, the team used cryo-EM, a relatively new technique that won the 2017 Nobel Prize in Chemistry, to determine the 3D shape of proteins, which are often thousands of times smaller than the width of a human hair. The technique works by blasting a stream of electrons at a flash-frozen sample to take many 2D images.
Seeing this structure allows us to understand how it works. It’s like taking apart a common household appliance to see how everything fits together.”
graduate student
For this study, He’s team captured hundreds of thousands of images of the Med-PIC complex. They then used computational methods to reconstruct a 3D image.
“Solving this complex was like assembling a puzzle,” Talyzina said. “Some of those subunits were already known from other experiments, but we had no idea how the pieces assembled together or interacted with each other. With our final structure, we were finally able to see this whole complex and understand its organization.”
The resulting image shows the Med-PIC complex as a flat, elongated structure, measuring 45 nanometers in length. The researchers also were surprised to discover that the Mediator moves relative to the rest of the complex, binding to RNA polymerase II at a hinge point.
“Mediator moves like a pendulum,” Abdella said. “Next, we want to understand what this flexibility means. We think it might have an impact on the activity of a key enzyme within the complex.”
The paper, “Structure of the human Mediator-bound transcription pre-initiation complex,” was supported by Northwestern’s Chemistry of Life Processes Institute, the Chicago Biomedical Consortium with support from the Searle Funds at The Chicago Community Trust, the American Cancer Society (award number IRG-15-173-21), the H Foundation (award number U54-CA193419) and the National Institutes of Health (award numbers R01-GM135651 and P01-CA092584).